|
the old AssemblyLIGNĬopyright © 2022 MacVector, Inc. Click here to learn more about MacVector Assembler vs. This means that when you create assembled contigs, you can immediately analyze the sequences using any of MacVector's DNA analysis or manipulation functions. Bidirectional reads are assembled by name into contigs, retaining the morphological identification and the. 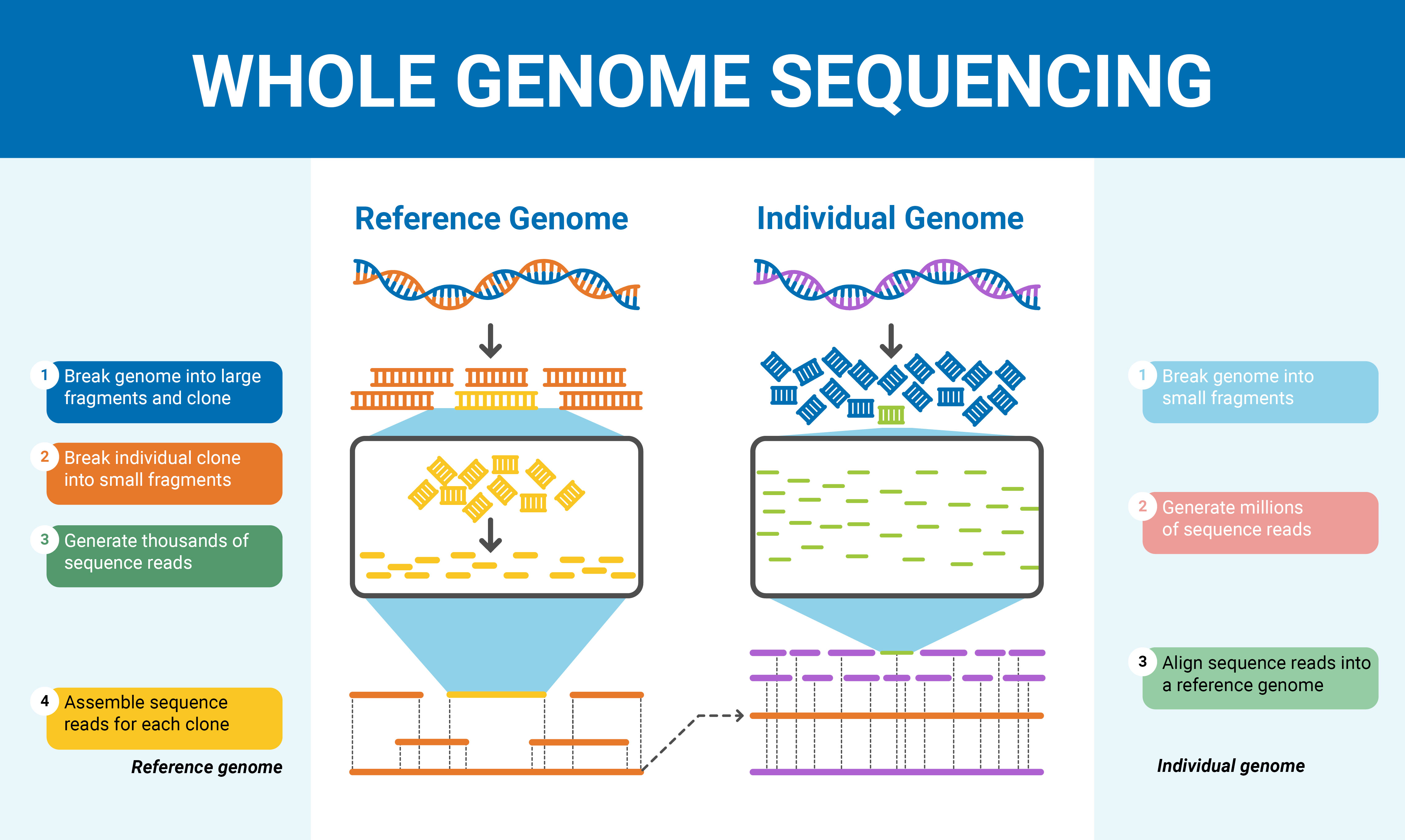
MacVector Assembler integrates tightly into MacVector so that they appear as a single application. We use GeneCodes Sequencher software for this. 
You can also generate genomic reference alignments with NGS data aligned using Bowtie which can also be used to examine expression analysis using RNAS-Seq data. MacVector Assembler integrates tightly into MacVector so that they appear as a single application. We demonstrate that short reads can be locally assembled into longer contigs using paired-end sequencing of restriction-site associated DNA (RAD-PE) fragments. 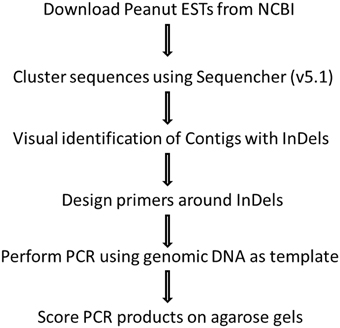
MacVector Assembler is an add-on module for MacVector that lets you assemble sequences using the phred/phrap/cross_match algorithms from the University of Washington, or the Velvet or SPAdes or Flye algorithms for next generation sequencing data. Despite the power of massively parallel sequencing platforms, a drawback is the short length of the sequence reads produced. Sequence Analysis Tools for Molecular Biologists The SnapGene 'Assemble Contigs' tool uses the CAP3 assembler to assemble reads into one or more contiguous assemblies.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
